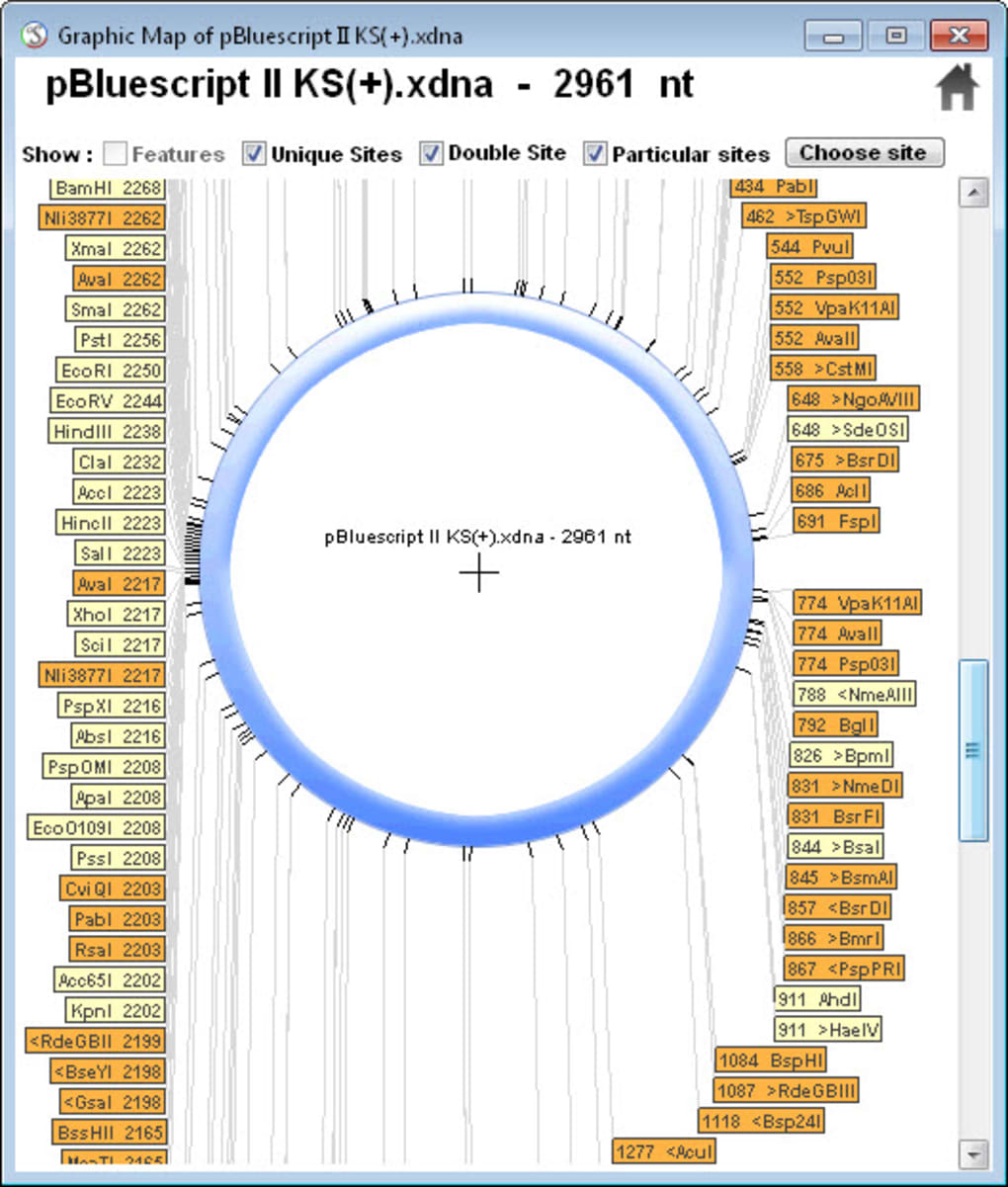
It says: CLC Genomics Workbench, for analyzing and visualizing next generation sequencing data, incorporates cutting-edge technology and algorithms, while also supporting and integrating with the rest of your typical NGS workflow. License: Freeware, Source code available as Xcode project It says: BioSAVE (the E probably comes from Viewer somewhere) is a program for visualising DNA or protein sequences and annotations thereof. It says: Design, analyze, and share sequence data in the cloud. Use Beacon Designer™ for optimal TaqMan® probes. Beacon Designer™ offers a comprehensive solution for mutation detection. You can BLAST search sequences and search for template structures from within the program. It says: Beacon Designer™ automates the design of real time primers and probes. You can use it for viewing DNA sequence, translation, creating graphic restriction maps, saves graphics as encapsulated postscript or scalable vector graphics, virtual restriction digest and a lot more. 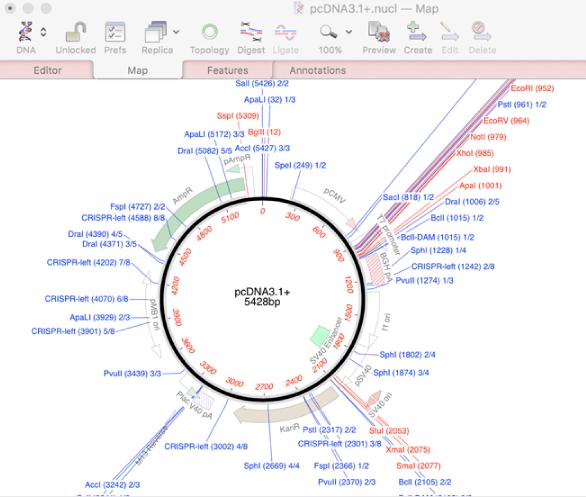
It can also be used as a tool for designing primers by evaluating candidates. It says: Amplify is a freeware Macintosh program for simulating and testing polymerase chain reactions (PCRs). It says: The award winning 4Peaks helps molecular biologists to visualize and edit their DNA sequence trace files. Getting Started with MacVector: An overview of primer design workflows in MacVector.Software for DNA, RNA and Cloning Software for DNA, RNA and Cloning.Melissa Caimano on HOW DO I video guides to common molecular biology workflows.admin on HOW DO I video guides to common molecular biology workflows.mariam abdelmalak on Major release details – Summary.Brian on Designing primers and documenting In-Fusion Cloning with MacVector.Chris on Designing primers and documenting In-Fusion Cloning with MacVector.How to call heterozygotes in trace files or Assembly Projects.MacVectorTip: How to Customize the Toolbars of MacVector windows.MacVectorTip: Selecting the sequence from a single restriction enzyme site to the end of a linear sequence.Sequence Assembly: What can Assembler do for my lab?.The window that opens contains a COPY of the starting sequence – now click on the Add Seqs button and select all the ABI/.ab1/SCF chromatogram files from your sequencing project directory to import them into the window In any event, open that sequence first, then choose Analyze | Align To Reference. It can even be the entire genome of an organism to which you want to align chromatograms from sequencing runs of mRNA or cDNA clones. Otherwise, you would normally start with a Reference sequence – it might be the predicted sequence of the PCR fragment you cloned, the starting sequence for a mutagenesis experiment or even a related sequence from another organism. I’m not going to discuss that here but there is a tutorial that you can download from this link. Admittedly, that is not always the case – if you are trying to determine the sequence of an unknown piece of DNA, then you need to use the add-on Assembler module. Typically, if you are aligning chromatograms it is because you are re-sequencing something. None of these algorithms will “flip” the chromatograms when required, but the strategies described below will do that. Note that you should NOT use the Multiple Sequence Alignment ( ClustalW, Muscle or T-Coffee) interface unless you are truly looking for the evolutionary relationships between the sequences and know that they are already all in the correct orientation. One common request we get is “I want to see my chromatograms/traces aligned so that I can edit the alignments”. The blog link discusses the 6 main alignment algorithms in MacVector and how to decide which is the most appropriate for accomplishing different tasks. I blogged about this a few years ago, but its something that still comes up on a regular basis.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
